AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
Back to Blog
Open file python paraview12/17/2023 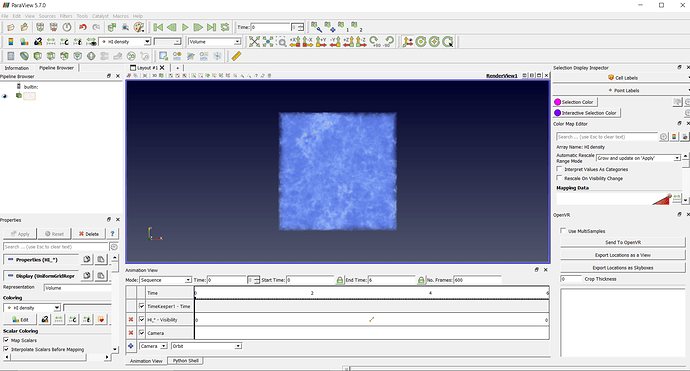 
This is the straight forward way to create an animation. By default, it will show the standard options to export an animation that contains all timesteps. Next, the dialog box Save Animation Options will appear. The individual frame number for each time step will be automatically added to your file name when the animation is created. Therefore, we typically choose the png-image format. We want to export each time step as an individual image. A new dialog box will allow you to specify the file name of the animation. To create an animation of time dependent data, select in the main menu File > Save Animation. How to create animations of time dependent data Here we commented (#) the original variable declaration and we added a new variable declaration that only contains the variable 'p'. For example, if you only want to visualize the pressure 'p', then you can modify the Python script as following: #casefoam.CellArrays = We recommend to modify the field CellArrays, so that it only contains the required variables. This is particularly important when reading large data sets or if you want to make animations with a large number of timesteps/images. This means, ParaView requires more memory to store all variables and longer time to read the variables from disk than necessary. If you did not adjust Cell Arrays, when opening your file in ParaView, the list may contain variables that you do not need for your actual visualization. The Python trace file contains a list of variables, that ParaView reads from disk: casefoam.CellArrays = Python trace file: How to adjust the list of variables You probably have to convert it into a different format to visualize your data with ParaView. It seems that ParaView does not support the collated file format so far. It reduces the number of files for parallel computations. ParaView loads your data, after you have clicked "Apply".Ĭollated file format: OpenFOAM also offers the collated file format, that has been introduced in OpenFOAM 7 and OpenFOAM v1712. For large datasets, it is recommended only to load the necessary variables. OpenFOAM: Decomposed CaseĬhoose your variables: Under Cell Arrays, you can select the variables that you want to load. The solution data will not be loaded, due to the wrong Case Type. If you try to read your decomposed data with Case Type = Reconstructed Case, then the field Cell Arrays will be empty. Depending if your data is saved as a decomposed case, or a reconstructed case, you have to choose the correct Case Type, when opening the file case.foam: Decomposed Case or Reconstructed Case. 
ParaView needs a *.foam file in your OpenFOAM case directory, which indicates that the data is saved in the OpenFOAM format, e.g. For more information on how to use ThinLinc, please see: Running graphical applications using ThinLinc When running the ParaView GUI, NSC recommends using ThinLinc to access Tetralith. The following table summarizes how to start ParaView with GPU or mesa rendering. How to start ParavView with mesa-software rendering is slightly different between ParaView/5.4.1-nsca-eb and ParaView/5.8.1-nsc1-bsidt. On a standard compute node without GPU, you have to use software rendering (mesa). On a Tetralith GPU node and on the login nodes, it is recommended to make use of the GPU. This depends on the ParaView version and the type of rendering, namely GPU accelerated rendering or software rendering (mesa). There are certain diffences how to start ParaView. This will add the paraview application to your search path. Load the paraview module corresponding to the version you want to use, e.g module load ParaView/5.4.1-nsca-eb
0 Comments
Read More
Leave a Reply. |
 RSS Feed
RSS Feed
